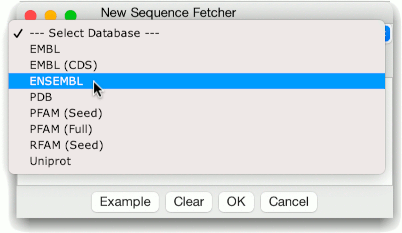
Two types of ENSEMBL source are provided. ENSEMBL queries the main ENSEMBL warehouse containing data for higher eukaryotes, and EnsemblGenomes, which queries Ensembl Pathogens, and other warehouses.
General Use
If you have a set of Ensembl
gene or transcript IDs, then you can retrieve them via the
sequence fetcher dialog opened after selecting the most appropriate
source (either 'ENSEMBL', or Ensembl Genomes). However, Jalview's
Ensembl client has a couple of additional capabilities:
Retrieving aligned transcripts for a genomic ID
If a single genomic identifier is entered in the Ensembl fetcher, Jalview will return all transcripts and products for the locus, and display them in a split view - complete with sequence variant annotation.
Retrieving orthologs for a gene ID
If a gene ID is entered (e.g. fox1), Jalview will resolve Ensembl
genomic identifiers for a predefined set of taxa (Mouse, Rat, Human,
Yeast in Jalview 2.10).
Ensembl Sequence
Features
Jalview 2.10 includes support for the
visualisation and transfer genomic and transcriptomic sequence
features onto protein product sequences. Retrieval of a genomic
locus results in a set of transcripts that are annotated with
nucleotide variant information and exonic regions. By default,
intronic regions will be hidden.
Variant information on
Protein Products
Jalview can translate genomic variant
annotation into protein sequence variant codes for variants
intersecting coding regions of a gene. To see this in action, use
the Calculate→Show cross-references menu to
view protein product sequences for the currently displayed (or
selected) sequences. The same menu allows you to recover Ensembl
exon, transcript and variant information when viewing UniProt
sequences.
Viewing more information about variant annotation
Variants are highlighted as red sequence features on the protein
sequence, with each one reporting all protein sequence variants
observed at that position as a result of the genomic variants.
Right-clicking a variant allows you to open the Ensembl Variants web
page for each variant, via the Link submenu.
Work in Progress !
In the next few releases,
we hope to improve and extend Jalview's support for working with
Ensembl. If you have any problems, questions or suggestions then
please get in contact with us via the Jalview discussion list.